Among-individual behavioural variation
Below are tutorials associated with “Avoiding the misuse of BLUP in behavioural ecology“ (Houslay & Wilson, 2017), published in the ISBE Behavioral Ecology journal. These tutorials are aimed at researchers interested in multivariate methods for modelling among-individual variation in labile traits (we focus on personality, behavioural syndromes, and individual variation in behavioural plasticity).
We have provided two versions of each tutorial – one using maximum likelihood via the R interface for VSNi commercial software ASReml, and the other using Bayesian methods via Jarrod Hadfield’s MCMCglmm R package.
The data sets for use with these tutorials are available on figshare:
https://figshare.com/projects/Avoiding_the_misuse_of_BLUP_in_behavioural_ecology/19426
If you have comments, feedback or further requests (including for reprints of the article), please email me. If you find these tutorials useful for your research, please cite Houslay & Wilson (2017).
Multivariate modelling for individual variation
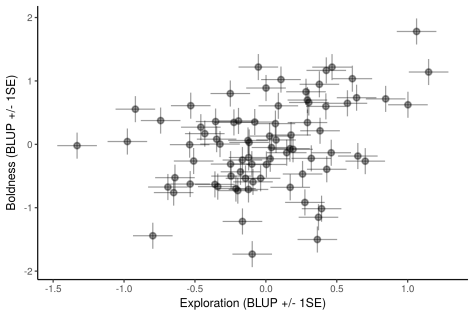
Here we demonstrate the use of multivariate models for directly testing among-individual correlations between ‘personality’ traits (measured repeatedly at the individual level), in addition to testing for associations between these traits and a single measure of fitness (measured once at the individual level).
ASReml-R: PDF
MCMCglmm: PDF
Data: figshare link
Multivariate modelling for individual plasticity variation

Here we show how to test for individual-by-environment interactions (also known as individual variation in slopes, or individual variation in plasticity) using random regression. We also show how to add a further response variable to this random regression, and test for an association between this variable and individual variation in intercepts/slopes. We then discuss problems with the interpretation of random regression models, and why a ‘character state’ approach might be beneficial. We show how to model individual variation in plasticity with character state models, and add the single fitness measure to this model to demonstrate this might be a more intuitive approach.
ASReml-R: PDF
MCMCglmm: PDF
Data: figshare link
Multivariate behavioural (co)variation
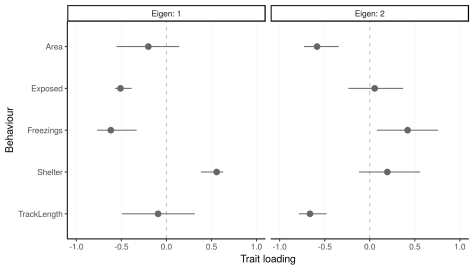
In this tutorial, we show how to isolate the among-individual (co)variance matrix I by applying a multivariate mixed model to a set of traits. We then examine eigen vectors of I to determine what the major axis of among-individual variation looks like. This is more relevant to personality / behavioural syndrome studies than the oft-used technique of applying univariate mixed models to principal components of multivariate data (since these PCs contain both among- and within-individual trait variation).
Note: while more in-depth tutorials on this topic are forthcoming, we have provided well-annotated R code in markdown format as part of the supplementary information for another recent paper, ‘Testing the stability of behavioural coping style across stress contexts in the Trinidadian guppy‘. This supplementary information uses ASreml-R to estimate the covariance matrices and does some fancy bootstrapping to estimate uncertainty. If you don’t have ASreml-R and are instead using MCMCglmm, you can use the posterior distributions from your MCMCglmm model object instead – I’ll try to get round to finishing up these additional tutorials, but until then hopefully this is a useful starting point!
The supplementary information and data can be accessed using the following links:
Supp info (annotated ASreml-R code in Appendix S1)
ASReml-R: coming soon…
MCMCglmm: coming soon…
Data: coming soon…
Plasticity and behavioural syndromes
David Mitchell and I co-authored a paper published in Behavioural Ecology, ‘Context-dependent trait covariances: how plasticity shapes behavioural syndromes‘, that focuses on the ways in which individual variation in plasticity can affect trait correlations. We lay this out in terms of behaviour, although this can be applied to other flexible traits. The supplements for this paper can be found here:
OSF|Supplements to ‘Context-dependent trait covariances’
These supplements focus largely on interpreting data and models, so we have not (as yet) supplied code. These models can be fit using guidance from the above tutorials, and there is useful code in brms in the supplements to this preprint of David’s:
Data visualisation
More tutorials coming soon…